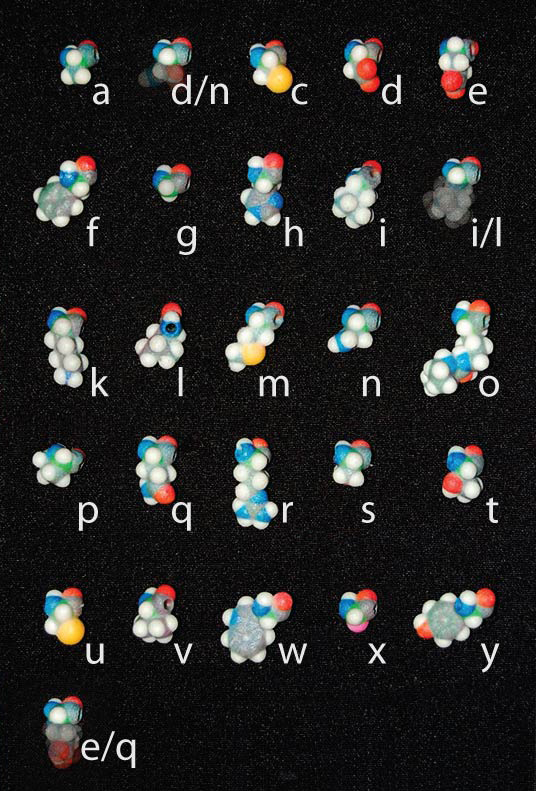
 Amino acids are known by there full names, a 3-letter abbreviation, and a single-letter abbreviation. For example, the amino acid, Proline, is recognized as Proline, Pro, or just P. We use the single-letter abbreviation to spell messages, names, or initials using our amino acid beads.
Amino acids are known by there full names, a 3-letter abbreviation, and a single-letter abbreviation. For example, the amino acid, Proline, is recognized as Proline, Pro, or just P. We use the single-letter abbreviation to spell messages, names, or initials using our amino acid beads.
The International Union of Pure and Applied Chemistry (IUPAC) and the International Union of Biochemistry and Molecular Biology (IUBMB) set the guidelines that uniquely identify each amino acid with their the single-letter code and three letter code.
Amino acids are the building blocks of proteins. Proteins are made of amino acids arranged in a specific order, like beads on a string. The sequence of these “beads” is known as a protein’s primary structure. The amino acids can form short chains called peptides, or long chains called polypeptides or proteins.
These amino acid chains can take up complex shapes as the protein folds, giving rise to a protein’s secondary, tertiary and even quaternary shape. The final shape, however, is dependent on the primary sequence, as each amino acid’s side chain lends different properties that influence it’s folding.
However, even if you know a protein’s primary structure, predicting the final shape is no easy feat. Many additional factors contribute to the final structure, such as pH, temperature, or presence of molecular chaperones. Large efforts have gone underway in the field of protein structure prediction. Scientists have even created a computer game, called Foldit, that calls upon users to help predict 3D structures of proteins.
Purchase Amino Acid Beads Here
Download Amino Acid Key
